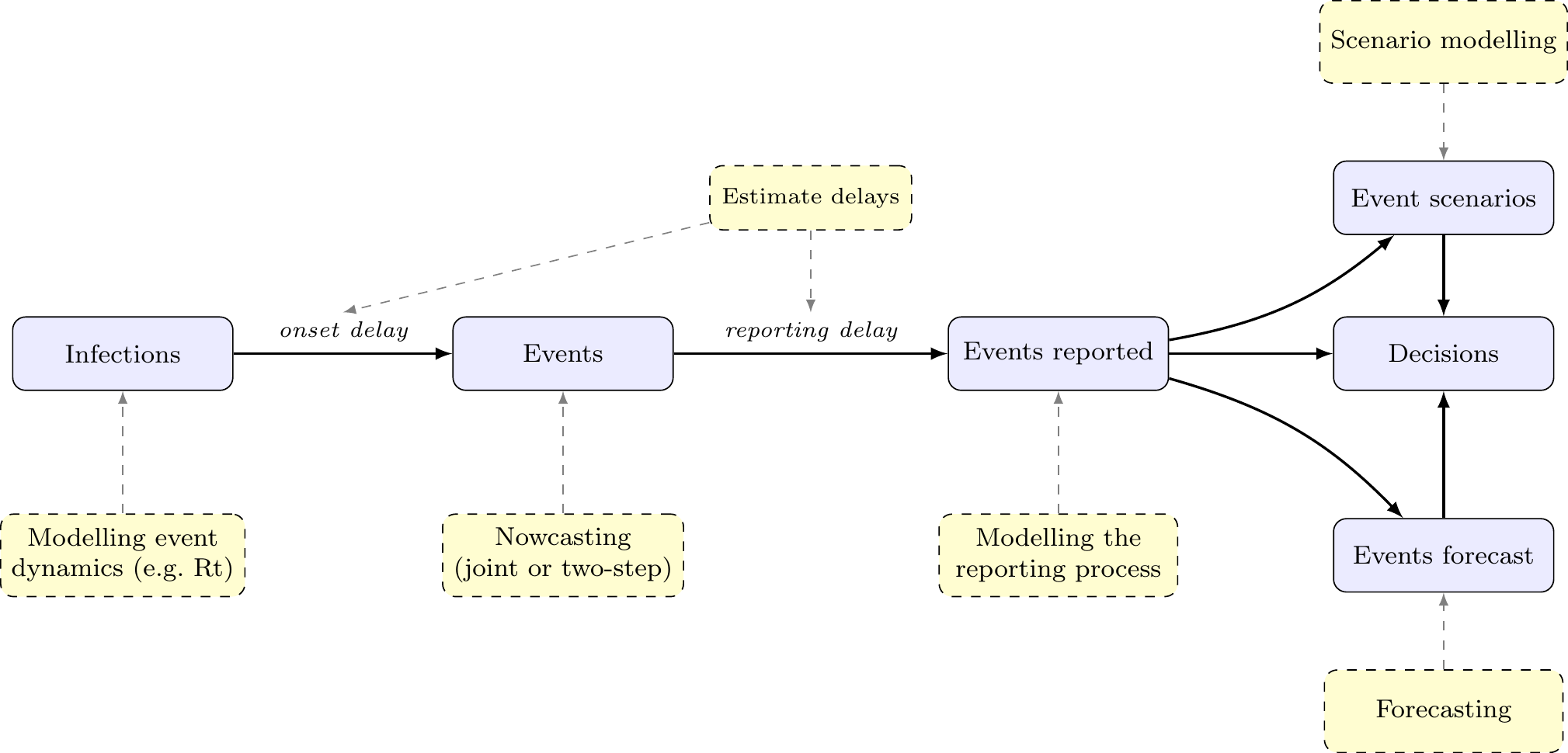
Modelling options for reporting delays
InSight Net nowcasting strike team
2026-04-26
From three examples to a methods view
You have just seen three concrete cases:
- Massachusetts (respiratory ED visits via NSSP)
- Washington State (case investigation, COVID-19)
- South Carolina (measles)
Different surveillance streams, different operational responses, but a similar underlying data picture.
Right truncation, and more
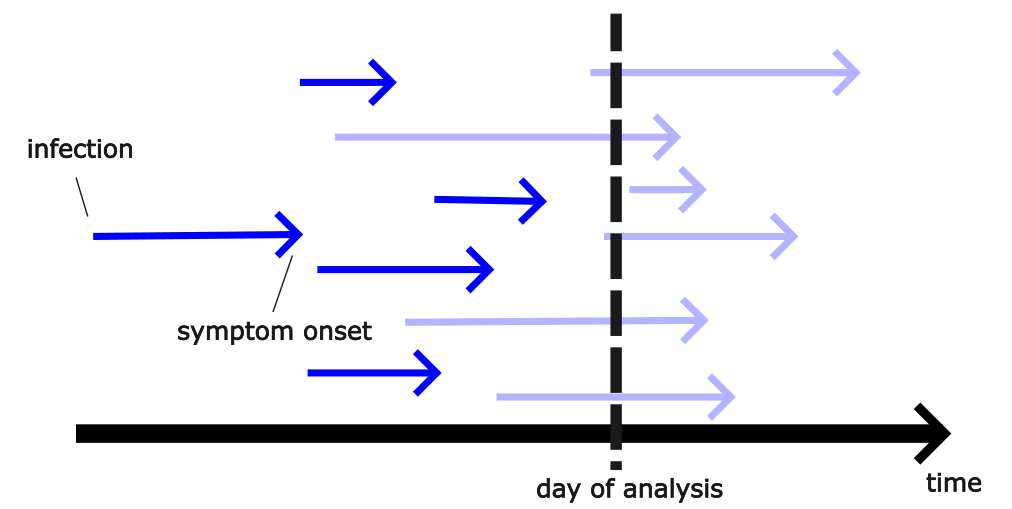
All three examples share right truncation: most events have happened, but some are not yet visible to us.
They also have other things going on — data revisions, sites coming and going, missing or messy fields — that need different kinds of methods.
Modelling for delays is broader than nowcasting alone
Reporting delays
Approaches
- Simple: pruning recent dates, caveats, switching to report date
- Individual-level delay estimation: fit a delay distribution to paired event/report records (line list); feed into other methods
- Multiplicative (chain ladder): scale partial counts by historical completion rates
- Regression: joint model of epidemic curve and delay surface, with covariates
- Generative: joint model of counts and delay distribution, with explicit structure (parametric delays, pooling, mechanism)
- Other statistical: machine learning, time-series methods
What you get: estimates of current levels and trends, usually with uncertainty.
Preliminary — we need your input.
Downwards data revisions
Approaches
- Nowcasting methods that do not assume monotonic (increasing) reporting, so they can absorb downward revisions (e.g. deduplication, reclassification)
- Direct modelling of the revision process across snapshots
What you get: estimates that do not flip when a later release revises earlier values down.
Preliminary — we need your input.
Site drop-in / drop-out
Approaches
- Pooling across sites, with explicit handling of which sites contributed at each time
- Reporting-population models that separate “what happened” from “who was reporting”
What you get: jurisdiction-wide estimates that stay coherent as sites join, leave, or report intermittently.
Preliminary — we need your input.
Other data quality
Approaches
- Pre-processing: deduplication, format harmonisation, strata cleaning
- Generative / Bayesian methods can be adapted to many of these — e.g.
epinowcastdoes missing-date imputation alongside the nowcast - More custom or ad-hoc work where no standard tool fits
What you get: principled handling of messy data, in the same model as the rest of the analysis where possible.
Preliminary — we need your input — please share your own approaches.
More detail in the guide’s modelling options section.
Our tools
Most of the methods we work on sit in the epinowcast community:
- epinowcast — modular Bayesian framework for nowcasting, Rt estimation, delay estimation, and forecasting from delayed data
- epidist, primarycensored, baselinenowcast — R packages for delay distribution estimation, primary event censored distributions, and simple baseline nowcasts
- EpiNow2 — R package for Bayesian Rt estimation, nowcasting, delay estimation, and forecasting with reporting delays
- The EpiAware org for the same ideas in Julia, including CensoredDistributions.jl
If any of these are relevant to your setting, please grab me after the session.
Our courses and seminar
Course — free, open-source, on nowcasting and forecasting:
- NFIDD — Nowcasting and forecasting of infectious disease dynamics, available online any time
- Taught in person at SISMID 2026 (Emory, July 2026): course site
Seminar series — talks from the wider community:
- epinowcast seminars — recurring online seminar series on nowcasting, forecasting, and real-time methods
Materials are open and reusable; happy to point you to specific sessions if a topic from today is useful for your team.
Up next: rapid talks on modelling
As you listen, things to note:
- What questions can it help answer?
- What data does it need?
- What does it give back?
- Where does it attach in the modelling workflow?
- What does it assume?
Modelling options for reporting delays